This guide covers everything you need to do before using AlgAware-IFCB for the first time. You will install R, RStudio, and the package, then configure the application for your cruise data.
1. Install R and RStudio
If you do not already have R and RStudio installed:
- Download R from https://cran.r-project.org/ and run the installer.
- Download RStudio Desktop (free) from https://posit.co/download/rstudio-desktop/ and run the installer.
You only need to do this once.
2. Install the AlgAware-IFCB package
Open RStudio and paste the following into the Console panel (bottom-left), then press Enter:
install.packages("remotes")
remotes::install_github("nodc-sweden/ifcb-algaware",
dependencies = TRUE,
ref = remotes::github_release())The dependencies = TRUE argument ensures all optional
packages are installed alongside the main package. Installation may take
a few minutes the first time.
Note for Linux users: Some system libraries must be installed before the R packages will compile. On Ubuntu/Debian, run the following in a terminal:
sudo apt-get install libhdf5-dev libudunits2-dev libgdal-dev libgeos-dev \ libproj-dev libmagick++-dev
3. Launch the application
The application opens in your default web browser. Leave the RStudio window open — closing it will stop the app.
4. Configure settings
Click the Settings tab in the left sidebar. You must fill in the paths and connection details before the app can load any data.
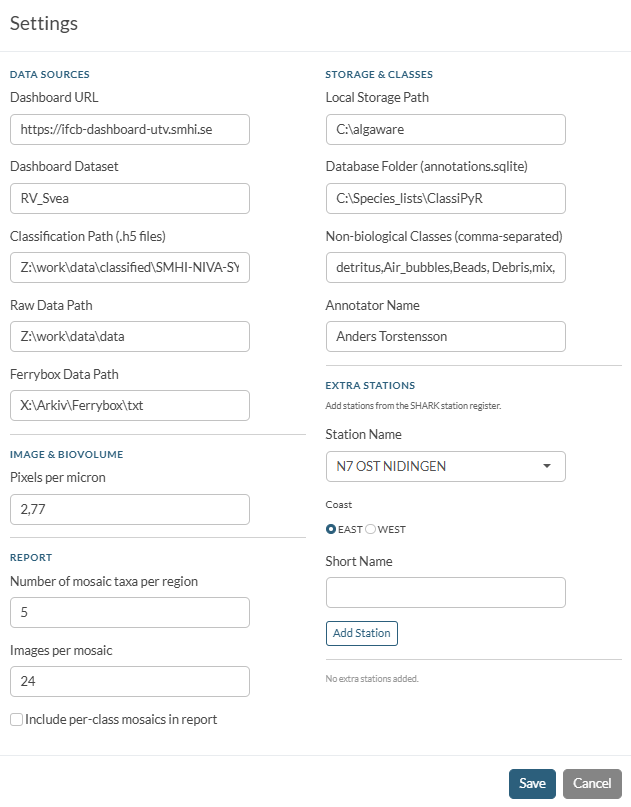
| Field | What to enter | Example |
|---|---|---|
| Dashboard URL | Base URL of the IFCB Dashboard instance | http://ifcb-dashboard-utv.smhi.se/ |
| Dashboard Dataset | Dataset name as it appears in the Dashboard | RV_Svea |
| Classification Path | Folder containing the HDF5 (.h5) classifier output
files |
C:/Users/name/classifications |
| FerryBox Data Path | Folder with FerryBox .csv files (optional) |
C:/Users/name/ferrybox |
| Local Storage Path | Where downloaded IFCB data will be cached on your computer | C:/Users/name/algaware_data |
| Database Folder | Where the annotations SQLite file will be stored | C:/Users/name/algaware_db |
| Non-biological Classes | Comma-separated class names excluded from all analyses | detritus,Air_bubbles,Beads,Debris,mix,mixed |
| Pixels per Micron | Calibration factor for the IFCB camera | 2.77 |
Click Save Settings when done. Settings are stored
in ~/.config/R/algaware/settings.json and reloaded
automatically next time you launch the app.
Adding monitoring stations from SHARK
If your cruise visited stations not yet in the built-in list, you can add them from the SHARK station register:
- In the Settings panel, click Load stations from SHARK.
- The app downloads the current station list from SHARK and merges it with the built-in AlgAware stations.
- Newly added stations will appear in all spatial-matching and reporting functions.
Note for package maintainers: Stations that should be permanently bundled in the package (rather than fetched at runtime) need to be added to
inst/stations/algaware_stations.tsv. See the section Bundled configuration files below.
5. Configure LLM API keys (optional)
AlgAware-IFCB can generate Swedish and English report text
automatically using OpenAI (GPT-4.1) or Google Gemini. This requires an
API key from the respective provider and the httr2 package
(installed with dependencies = TRUE).
Set up the key in RStudio
Open your .Renviron file by running:
usethis::edit_r_environ()Add one of the following lines (replace the placeholder with your actual key):
OPENAI_API_KEY=sk-proj-xxxxxxxxxxxxxxxxxxxxxxxxor
GEMINI_API_KEY=AIzaxxxxxxxxxxxxxxxxxxxxxxxxxxxxxxxSave the file, then restart R (Session → Restart R in RStudio). The key will be available to AlgAware-IFCB every time you launch the app.
If both keys are set, OpenAI is used by default. Override the model by adding:
OPENAI_MODEL=gpt-4.1
GEMINI_MODEL=gemini-2.5-flash-liteSecurity: Never share your
.Renvironfile or paste your API key into chat or email.
Bundled configuration files
AlgAware-IFCB ships several data files in inst/extdata/
and inst/stations/ that control how stations, taxa, and
plots are handled. A package maintainer can edit these files and rebuild
the package to change default behaviour without touching any R code.
inst/stations/algaware_stations.tsv
Defines the 12 AlgAware monitoring stations used for spatial bin-matching and the chlorophyll map. Each row is one station:
| Column | Description |
|---|---|
STATION_NAME |
Full canonical name (must match SHARK) |
COAST |
EAST (Baltic Sea) or WEST (West
Coast) |
STATION_NAME_SHORT |
Short label shown in plots and reports |
To add a new AlgAware station, append a row and rebuild the package
with devtools::install().
inst/extdata/standard_stations.yaml
Lists all standard Swedish monitoring stations (not just AlgAware) used for CTD profile matching and the regional chlorophyll time series. Stations are grouped by region:
standard_stations:
station_list: ["FLADEN", "ANHOLT E", ...]
FLADEN: "The Kattegat and The Sound"
ANHOLT E: "The Kattegat and The Sound"To add a station to a CTD region, add its name to
station_list and add a
StationName: "Region name" line. Existing region names are:
The Skagerrak, The Kattegat and The Sound,
The Southern Baltic, The Western Baltic,
The Eastern Baltic.
inst/extdata/station_mapper.txt
A tab-delimited synonym table mapping raw station name variants (as
they appear in CNV file headers or LIMS exports) to the canonical
station names used in standard_stations.yaml. Columns:
synonym, statn.
If a new station’s raw name in CNV files differs from its canonical name, add a row here. Names are matched case-insensitively, with prefix matching as a fallback.
inst/extdata/taxa_lookup.csv
Maps classifier class names to display names, WoRMS AphiaIDs, HAB flags, and italic/non-italic formatting. Edit this file to add new taxa, rename display labels, or update HAB status.